Purpose:
To make the Red Florescent Protein (RFP) from jelly fish in our bacteria.
To learn about the steps involved in genetic engineering.
To make the Red Florescent Protein (RFP) from jelly fish in our bacteria.
To learn about the steps involved in genetic engineering.
Materials and Procedure:
The materials and procedure can be found on the links below
2a. The materials and the procedure can be found in the Amgen lab manual, part 2a.
4a. The materials and the procedure can be found in the Amgen lab manual, part 4a.
5a. The materials and the procedure can be found in the Amgen lab manual, part 5a.
6. The materials and the procedure can be found in the Amgen lab manual, part 6.
The materials and procedure can be found on the links below
2a. The materials and the procedure can be found in the Amgen lab manual, part 2a.
4a. The materials and the procedure can be found in the Amgen lab manual, part 4a.
5a. The materials and the procedure can be found in the Amgen lab manual, part 5a.
6. The materials and the procedure can be found in the Amgen lab manual, part 6.
|
|
|
Experimental Overview:
Part 2a: We first needed to verify the plasmid by using restriction digest. Then we cut the plasmid using BamHI and Hind III to cut out the RFP-ara.
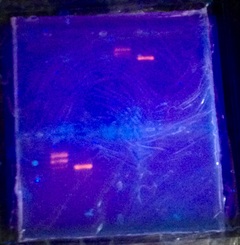
Part 4a: In this part we verified the plasmid digest by electrophoresis.
Part 5a: Next we transformed the bacteria with the recombinant plasmid.
Part 6: Finally we purified the RFP using chromatography.
Part 2a: We first needed to verify the plasmid by using restriction digest. Then we cut the plasmid using BamHI and Hind III to cut out the RFP-ara.
Part 4a: In this part we verified the plasmid digest by electrophoresis.
Part 5a: Next we transformed the bacteria with the recombinant plasmid.
Part 6: Finally we purified the RFP using chromatography.
Results:
All lab Questions can be found in the lab manual links above. All the answers to the questions can be found below.
Before the 2a Lab:
1. There are two fragments produced, the RFP with PBAD and Ara-C with ori with Amp-R. The RFP and PBAD equal 807 BP, and Ara-C, ori, and Amp-R equal 4495 BP.
2. The RFP gene and Ara-C are needed.
3. The selectable marker allows you to separate the bacteria that doesn't have the desired gene.
Part 2a Questions:
1. Ori- origin of replication; RFP- gene of interest; Amp-R - antibiotic resistance and selective marker; Ara-C - binds to promoter
2. The resistance enzymes are used to cut up unwanted bacteria DNA.
3. The bacteria keeps antibiotic resistant genes in order to survive so in modern medicine new stronger antibodies are created to kill bacteria and old ones no longer work.
4. Central dogma is the same in all organism.
5. Take half of the mixture and put each half in a petri dish. Put Amp in one dish and Kan in the other to kill Kan,in amp dish, and kill Amp,in Kan dish. This separates both Amp and Kan from each other.
Part 4a Questions;
1. The point in making ampicillin resistant bacteria is so the cells with Amp-R genes will live.
2. The pARA-R plasmid will only turn on the promoter if the bacterial cells are given arabinose. Without arabinose, then they will not turn red.
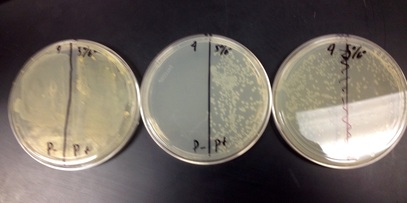
3. P- and P+ will grow on the LB plate, only P+ will grow on the LB/Amp plate, and a small amount of bacteria will grow on the LB/Amp/Ara plate.
All lab Questions can be found in the lab manual links above. All the answers to the questions can be found below.
Before the 2a Lab:
1. There are two fragments produced, the RFP with PBAD and Ara-C with ori with Amp-R. The RFP and PBAD equal 807 BP, and Ara-C, ori, and Amp-R equal 4495 BP.
2. The RFP gene and Ara-C are needed.
3. The selectable marker allows you to separate the bacteria that doesn't have the desired gene.
Part 2a Questions:
1. Ori- origin of replication; RFP- gene of interest; Amp-R - antibiotic resistance and selective marker; Ara-C - binds to promoter
2. The resistance enzymes are used to cut up unwanted bacteria DNA.
3. The bacteria keeps antibiotic resistant genes in order to survive so in modern medicine new stronger antibodies are created to kill bacteria and old ones no longer work.
4. Central dogma is the same in all organism.
5. Take half of the mixture and put each half in a petri dish. Put Amp in one dish and Kan in the other to kill Kan,in amp dish, and kill Amp,in Kan dish. This separates both Amp and Kan from each other.
Part 4a Questions;
1. The point in making ampicillin resistant bacteria is so the cells with Amp-R genes will live.
2. The pARA-R plasmid will only turn on the promoter if the bacterial cells are given arabinose. Without arabinose, then they will not turn red.
3. P- and P+ will grow on the LB plate, only P+ will grow on the LB/Amp plate, and a small amount of bacteria will grow on the LB/Amp/Ara plate.
Part 5a Questions:
1. Yes, our results were correct because we had growth on the LB plate for P+ and P-, the LB/Amp plate only grew on the positive side, and the LB/Amp/Ara plate had growth.
2. Their were no red colonies apparent most likely from temperature or not enough time staying in the incubator.
3. The RFP can only appear on the LB/Amp/Ara plate because RFP needs arabinose to be expressed.
4. The more copies apparent, then the more likely the promoter will be on and visible.
5. The RFP gene is shown from a trait through transcription by DNA --> mRNA --> Protien (process of central dogma).
6. Since central dogma is the same in all organisms, bacteria can replicate human genes.
1. Yes, our results were correct because we had growth on the LB plate for P+ and P-, the LB/Amp plate only grew on the positive side, and the LB/Amp/Ara plate had growth.
2. Their were no red colonies apparent most likely from temperature or not enough time staying in the incubator.
3. The RFP can only appear on the LB/Amp/Ara plate because RFP needs arabinose to be expressed.
4. The more copies apparent, then the more likely the promoter will be on and visible.
5. The RFP gene is shown from a trait through transcription by DNA --> mRNA --> Protien (process of central dogma).
6. Since central dogma is the same in all organisms, bacteria can replicate human genes.
Part 6(a,b) Questions:
1a. The red florescent protein (RFP) can be seen separated because the cells are red and florescent.
2a. The supernatant was fairly clear and the pellet had a pink or light red tint.
1b. BB (Binding Buffer) - makes amino acid and protein bind to the resin beads.
WB (Wash Buffer) - removes the proteins not bound to the resin beads
EB (Elution Buffer) - takes protein off resin beads
CEB (Column Equilibration Buffer) - stores resin beads
2b. This time the supernatant was more of a see through pink and the pellet was more of a dark pink/red color.
1a. The red florescent protein (RFP) can be seen separated because the cells are red and florescent.
2a. The supernatant was fairly clear and the pellet had a pink or light red tint.
1b. BB (Binding Buffer) - makes amino acid and protein bind to the resin beads.
WB (Wash Buffer) - removes the proteins not bound to the resin beads
EB (Elution Buffer) - takes protein off resin beads
CEB (Column Equilibration Buffer) - stores resin beads
2b. This time the supernatant was more of a see through pink and the pellet was more of a dark pink/red color.

Data Analysis/conclusion:
In the end we were able to create the red fluorescent protein from the bacteria we had. We were able to do this by isolating the RFP gene, testing it through electrophoresis, transferring it to the bacteria to be grown, and finally purified the RFP through chromatography. It may have taken a while but we ended up making the red fluorescent protein and discovered how bioengineering works. After running our purified protein through a vertical gel, we were able to find it and match it to a protein on the DNA ladder. The protein was pure because it appeared to be only one DNA band. The protein wasn't extremely concentrated since it wasn't very bold but was still visible. Our protein was visible around the 15-25 mark on the DNA ladder. Our protein matched fairly well to the desired protein level on the DNA ladder. This means our whole lab worked even though it may be hard to see.
In the end we were able to create the red fluorescent protein from the bacteria we had. We were able to do this by isolating the RFP gene, testing it through electrophoresis, transferring it to the bacteria to be grown, and finally purified the RFP through chromatography. It may have taken a while but we ended up making the red fluorescent protein and discovered how bioengineering works. After running our purified protein through a vertical gel, we were able to find it and match it to a protein on the DNA ladder. The protein was pure because it appeared to be only one DNA band. The protein wasn't extremely concentrated since it wasn't very bold but was still visible. Our protein was visible around the 15-25 mark on the DNA ladder. Our protein matched fairly well to the desired protein level on the DNA ladder. This means our whole lab worked even though it may be hard to see.
Reflection:
This lab was very interesting, long, and precise. I liked how making the red fluorescent protein could work with any gene since bacteria and humans make proteins the same way. I found this very interesting and also that bacteria genes can easily be replaced. Our group worked together fairly well and we finished everything at a reasonable pace. We almost always were able to answer each others questions and when we couldn't, someone stepped up to ask it. If we were to do this lab again we would read directions more carefully to avoid rereading them. In the end we thought we messed up but our protein came out in the right place unlike many groups. Overall this lab went well and we all worked together well.
This lab was very interesting, long, and precise. I liked how making the red fluorescent protein could work with any gene since bacteria and humans make proteins the same way. I found this very interesting and also that bacteria genes can easily be replaced. Our group worked together fairly well and we finished everything at a reasonable pace. We almost always were able to answer each others questions and when we couldn't, someone stepped up to ask it. If we were to do this lab again we would read directions more carefully to avoid rereading them. In the end we thought we messed up but our protein came out in the right place unlike many groups. Overall this lab went well and we all worked together well.